For Week 50 of the R4datascience project, I used the provided diseases data mentioned in this SimplyStats blog post. The dataset contains state level data of vaccine preventable infectious diseases over the span of ~70 years.
Packages
#packages
library(tidyverse)
library(gganimate)
library(ggplot2)
library(grid)
library(sf)
library(usmap)
library(scales)
library(Hmisc)
Read in the diseases file
diseases<-read.csv(file="~/GitHub/TidyTuesday/data/2019/2019-12-10/diseases.csv")
View the dataset
diseases %>%dim()
## [1] 18870 6
The diseases dataset contains 18870 rows and 6 columns
diseases %>% colnames()
## [1] "disease" "state" "year" "weeks_reporting"
## [5] "count" "population"
The diseases dataset contains columns for: disease, state, year, weeks_reporting, count, state population
head(diseases)
## disease state year weeks_reporting count population
## 1 Hepatitis A Alabama 1966 50 321 3345787
## 2 Hepatitis A Alabama 1967 49 291 3364130
## 3 Hepatitis A Alabama 1968 52 314 3386068
## 4 Hepatitis A Alabama 1969 49 380 3412450
## 5 Hepatitis A Alabama 1970 51 413 3444165
## 6 Hepatitis A Alabama 1971 51 378 3481798
The first 6 rows of the diseases dataset
diseases %>% summary()
## disease state year weeks_reporting
## Hepatitis A:2346 Alabama : 370 Min. :1928 Min. : 0.00
## Measles :3876 Alaska : 370 1st Qu.:1956 1st Qu.:14.00
## Mumps :1836 Arizona : 370 Median :1977 Median :44.00
## Pertussis :3774 Arkansas : 370 Mean :1974 Mean :33.28
## Polio :3774 California: 370 3rd Qu.:1992 3rd Qu.:50.00
## Rubella :1938 Colorado : 370 Max. :2011 Max. :52.00
## Smallpox :1326 (Other) :16650
## count population
## Min. : 0.0 Min. : 86853
## 1st Qu.: 1.0 1st Qu.: 1046542
## Median : 47.0 Median : 2824918
## Mean : 1367.5 Mean : 4242911
## 3rd Qu.: 440.8 3rd Qu.: 5153640
## Max. :132342.0 Max. :37607525
## NA's :204
unique (diseases$disease)
## [1] Hepatitis A Measles Mumps Pertussis Polio Rubella
## [7] Smallpox
## Levels: Hepatitis A Measles Mumps Pertussis Polio Rubella Smallpox
The diseases listed in the diseases dataset
diseases %>% select("count") %>% by(diseases$disease,sum)
## diseases$disease: Hepatitis A
## [1] 976362
## ------------------------------------------------------------
## diseases$disease: Measles
## [1] 18670996
## ------------------------------------------------------------
## diseases$disease: Mumps
## [1] 832110
## ------------------------------------------------------------
## diseases$disease: Pertussis
## [1] 2329020
## ------------------------------------------------------------
## diseases$disease: Polio
## [1] 2329020
## ------------------------------------------------------------
## diseases$disease: Rubella
## [1] 434966
## ------------------------------------------------------------
## diseases$disease: Smallpox
## [1] 232828
I summarized the cases of each disease by counts to get an idea of which one had the most cases in all of US over the span of the 76 years in the dataset. As anticipated, measles had the greatest number, with 18,670,996 cases in the United States from 1928 to 2003.
Calculate incidence rate
To compare the number of cases between states, I mutated the dataset to add an incidence rate that takes into account the case count over state population by 100,000 people for each state per year.
#Calculate incidence
diseases<- diseases %>%
mutate(incidence=(count/population*100000*(ifelse(weeks_reporting==0,0,52/weeks_reporting))))
Filter measles from the dataset
Because measles had the most cases of all diseases in the dataset, I decided to filter the dataset and explore measles for my visualization
#filter by measles/summary
diseases %>% filter(disease=="Measles") %>% summary()
## disease state year weeks_reporting
## Hepatitis A: 0 Alabama : 76 Min. :1928 Min. : 0.00
## Measles :3876 Alaska : 76 1st Qu.:1947 1st Qu.:32.00
## Mumps : 0 Arizona : 76 Median :1966 Median :47.00
## Pertussis : 0 Arkansas : 76 Mean :1966 Mean :37.45
## Polio : 0 California: 76 3rd Qu.:1984 3rd Qu.:50.00
## Rubella : 0 Colorado : 76 Max. :2003 Max. :52.00
## Smallpox : 0 (Other) :3420
## count population incidence
## Min. : 0 Min. : 86853 Min. : 0.0000
## 1st Qu.: 8 1st Qu.: 966309 1st Qu.: 0.3458
## Median : 543 Median : 2590843 Median : 27.7595
## Mean : 4817 Mean : 3858493 Mean : 166.6918
## 3rd Qu.: 3986 3rd Qu.: 4696902 3rd Qu.: 226.1823
## Max. :132342 Max. :34861711 Max. :2964.4269
## NA's :64 NA's :64
#create a measles dataset
measles<-diseases %>% filter(disease=="Measles")
measles<-measles %>% mutate (incidence_rounded = round(incidence, digits=1))
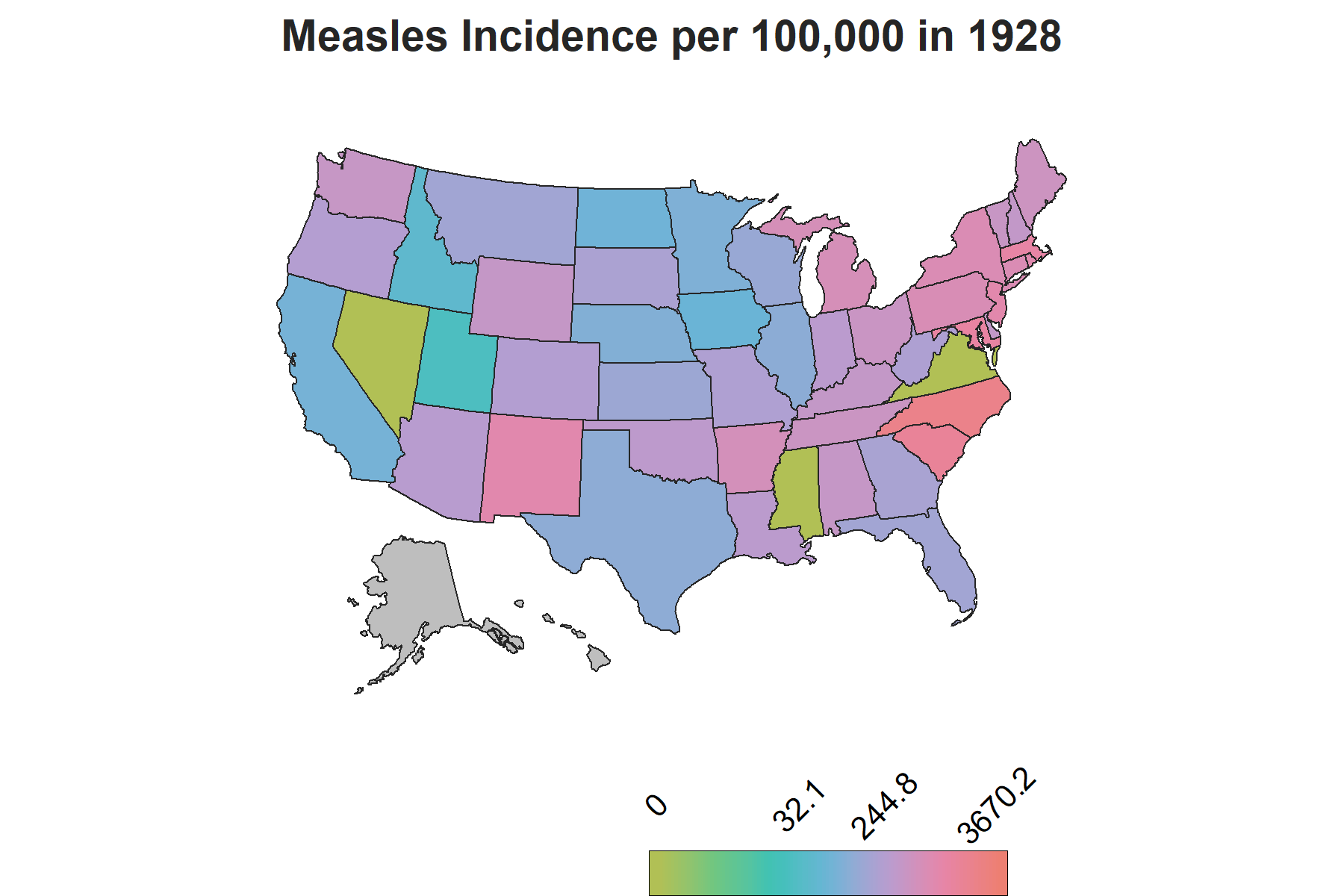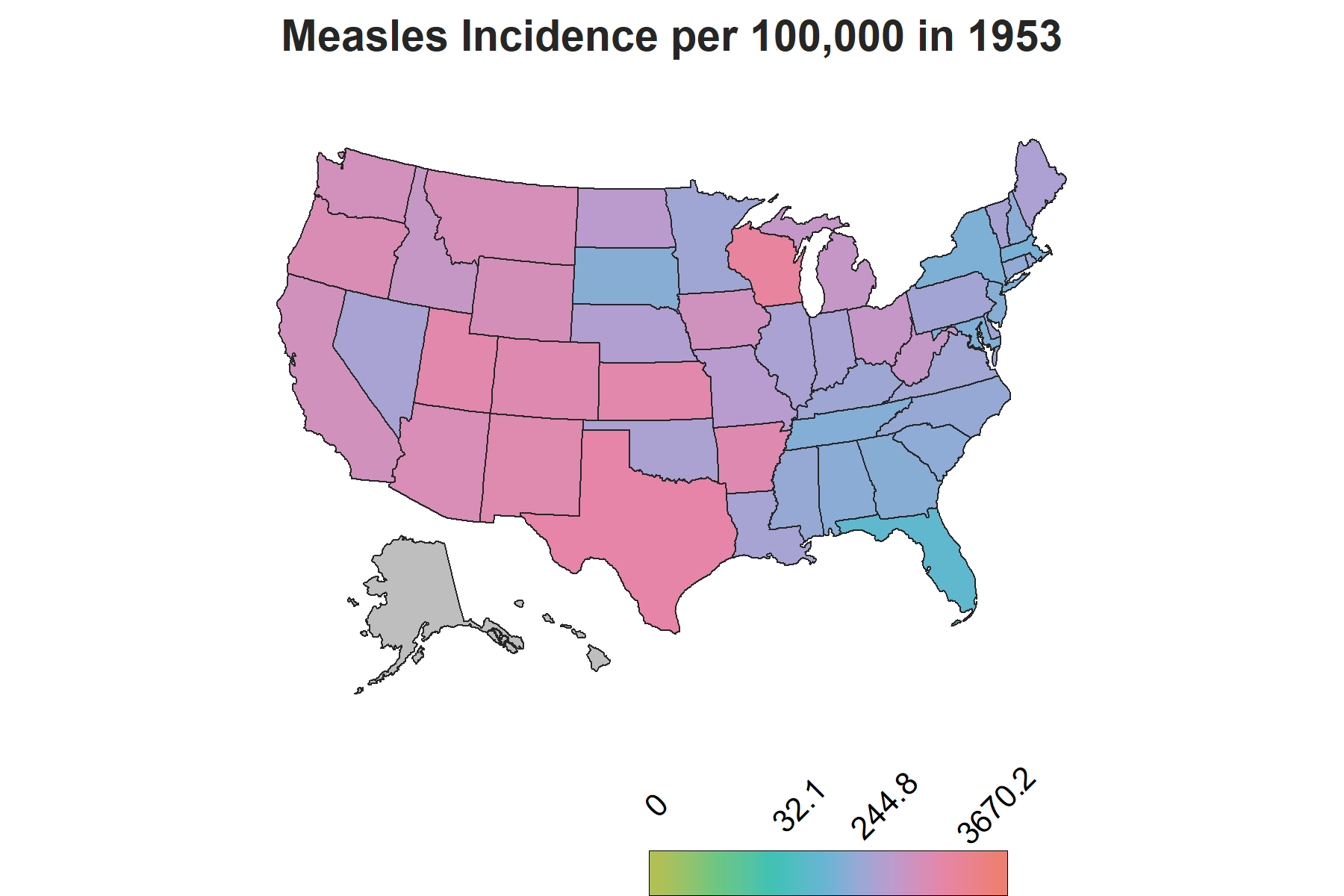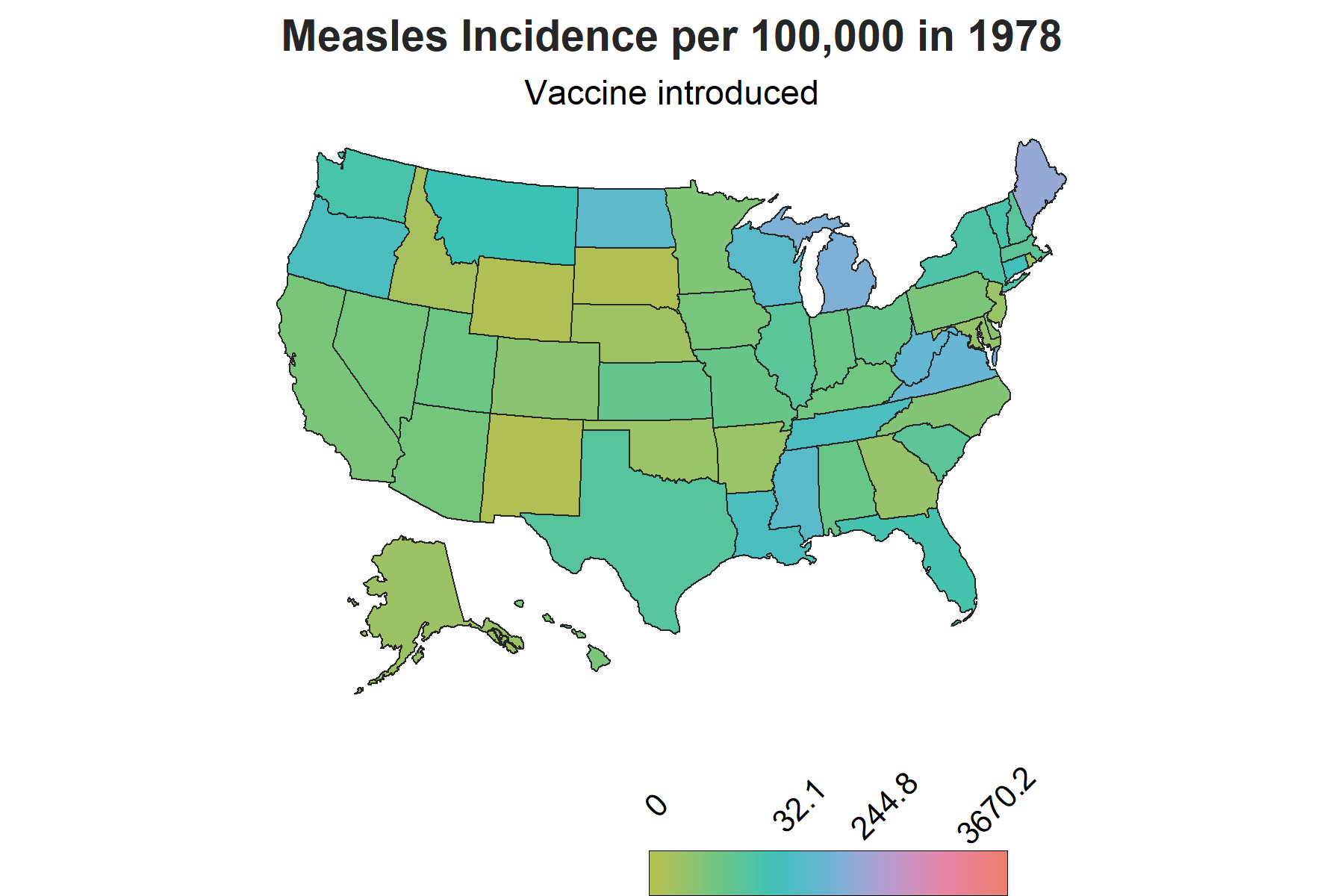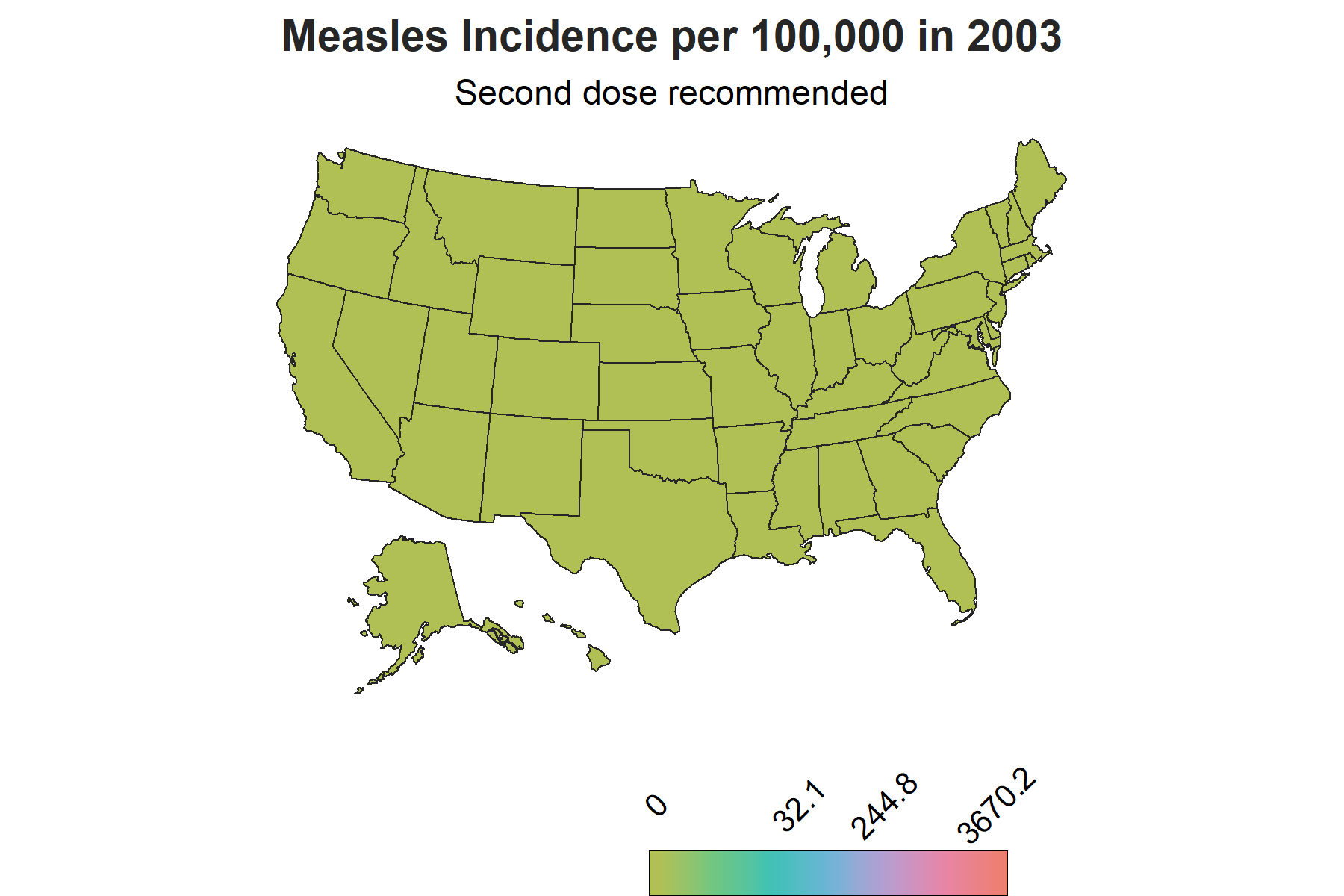
Since the dataset has data for each state by year, I decided it would be ideal to use a chloropleth map with gganimate to show how the incidence rate varies by each state over the years. The usmaps package provides one of many options to display state level data on a US map, which can also be done using the spData package, however with additional steps to include Alaska and Hawaii.
Create a FIPS column using fips() from usmaps package
The usmaps package has a fips() function that outputs the FIPs code for a state name string input. To have the state level data plotted onto the map, a FIPs column corresponding to each state needs to be created.
#Create a FIPS column using the fips() function from usmap
measles$fips<-fips(measles$state)
Create a new column to classify year by vaccine availability-will be used for subtitle in ggplot
There is another important piece of information to account for in the visualization to make sense of the changes in trends: vaccine introduction. The measles vaccine was introduced in 1963 in the United States with a second dose recomended by 1989. To show this on the map, I created a factor variable corresponding each year to three phases of vaccine availability: the time frame when there was no vaccine, the time frame when the vaccine was introduced and the years onwards when the second dose was reccomended. This column will be used as a the subtitle on the map to demonstrate the trends relative to vaccine availability over the course of the years.
#Create a column to classify year by vaccine availability
measles$vaccine<-ifelse((measles$year>=1963) & (measles$year<1989),2,
ifelse((measles$year>=1989),3,
ifelse((measles$year<1963), 1,1)))
measles$vaccine<-recode(measles$vaccine, '1'="",
'2'="Vaccine introduced",
'3'="Second dose recommended")
Creating quantiles for gradient breaks
measles %>% select(incidence) %>% summary()
## incidence
## Min. : 0.0000
## 1st Qu.: 0.3458
## Median : 27.7595
## Mean : 166.6918
## 3rd Qu.: 226.1823
## Max. :2964.4269
## NA's :64
If you take a look at the summary stats for the incidence in the measles data, you can see how variability between the rates would unevenly polarize rates above and below the third quartile, which would obscure interpretation of the rates on a gradient scale. Instead of using the default scale in scale_color_gradient() in the ggplot2 package which creates breaks by evenly spacing the intervals, quartiles are a better quantification of breaks in the dataset. a breaks vector is created to be read into the’breaks’ argument of the scale_color_gradient() function in the plot.
#Quantile breaks for gradient scale
breaks<- quantile(measles$incidence_rounded, probs=seq(0,1,.25), na.rm=TRUE) %>% unname() %>% round(digits=1)
Display
A few other things I added here in my code is a vector for a manual gradient color scheme and the resolution for the plot .gif file.
#Color palette
gradient<-c("#B1C055","#6CC682","#39C2B6","#6BB4D7","#B69DD1","#E685A8","#ED7F72")
#Set these image quality options
options(gganimate.dev_args = list(width = 6, height = 4, units = 'in', res=300))
Create the plot
#Create the plot
measles_plot<-plot_usmap(data=measles, color="#262626", size=.3, values="incidence_rounded")+
theme_void()+
scale_fill_gradientn(colors = gradient, trans="pseudo_log",
#The pseudo_log allows for log transformation even though 0 is in the dataset
na.value="grey",limits=c(min(breaks), max(breaks)),breaks=breaks[c(1,3:5)], labels=breaks[c(1,3:5)])+
#only including min, 50th, 75th and max values
guides(fill = guide_colorbar(title="",
frame.colour = "black",
label.position="top",
barwidth = 8,
barheight = 1,
ticks=FALSE,
keywidth=15,
label.hjust = 0.5,
label.vjust = 0.3,
label.theme = element_text(angle = 45, size=10)))+
labs(title = "Measles Incidence per 100,000 in {frame_time}",
subtitle="{unique(measles$vaccine[measles$year==(frame_time)])}")+
#Allows for vaccine column to display as a subtitle relative to plot animation
theme(legend.position="bottom", legend.justification=c(.8,0),
plot.title=element_text(face="bold", size=14, color="#262626",hjust=.5),
plot.subtitle=element_text(hjust=.5))+
transition_time(year)
Save
anim<-animate(measles_plot, nframes=76, fps=1)
#76 frames because 76 years
anim_save("measlesmap.gif", anim)
#save the map
The Map